Table of contents
Annotation Management
Experimental data is annotated by assigning extracts and therby samples to resources. A sample contains general information about the biological source while an extract represents an extraction of that source which actually is used for the experiment or measurement. Each sample and extracts consist of a number of attributes (annotations) each of which has its own vocabulary. These vocabularies can be extended by the users. Newly added annotations are then reviewed and released by FGCZ experts. There are two functionalities provided by B-Fabric while extending the vocabularies. First, any eligible user other than FGCZ experts is able to be defined as an annotation manager so that the review and release process can be done in a broader application of B-Fabric. Second, users are allowed to select annotations from standard vocabulary repositories and to import them into B-Fabric.Annotation Managers
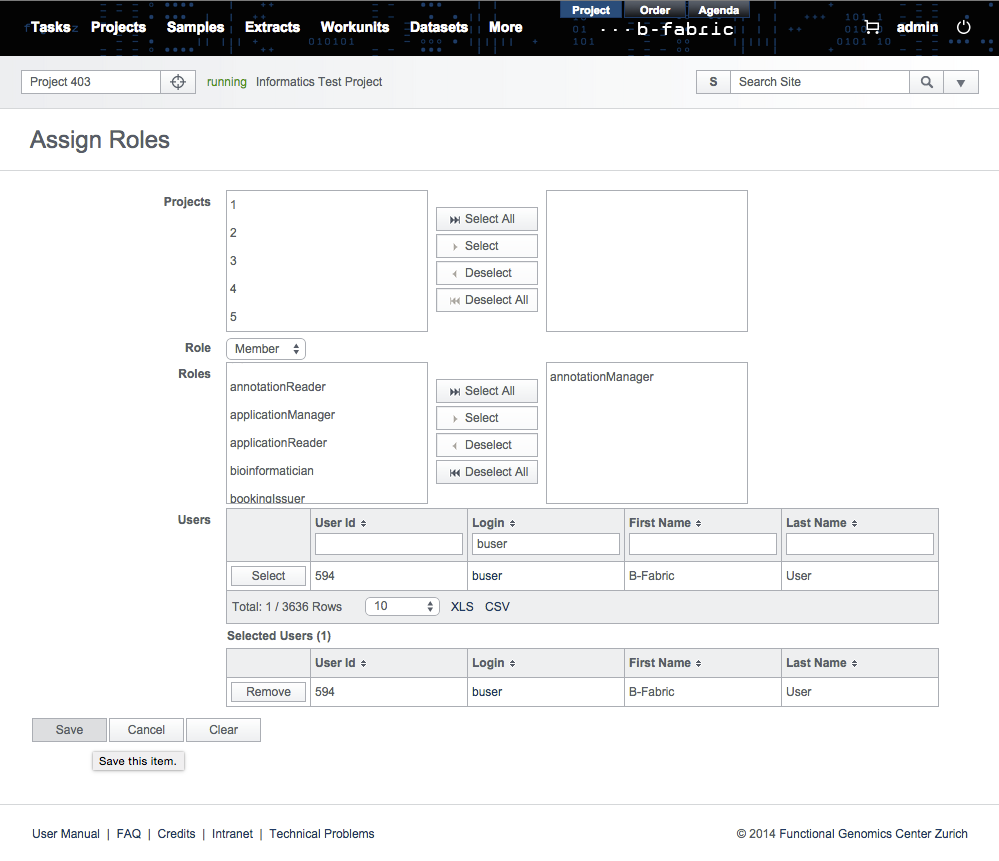
To become an annotation manager, a user must have the role annotationManager. This role can be granted by a user with the admin role to any user via "edit user" screen as below.Once the user is granted with the annotationManager role, he can review and release the newly added annotations. This task is achieved through "edit annotation" screen. The reviewer typically checks if the annotation is spelled correctly and proper for the selected type. He also checks the description of the annotation and revises it if necessary. When all checks are complete, the annotation is released for use by saving the annotation after having the released checkbox field ticked.
Importing Annotations
There are two approaches for importing annotations into B-Fabric. First, annotations can be imported from a specially formatted local file. This approach is convenient for batch import of several annotations at once. Second, annotations can be fetched from external repositories. This approach provides search functionality and is convenient for batch import as well.Import Annotations from a Local File
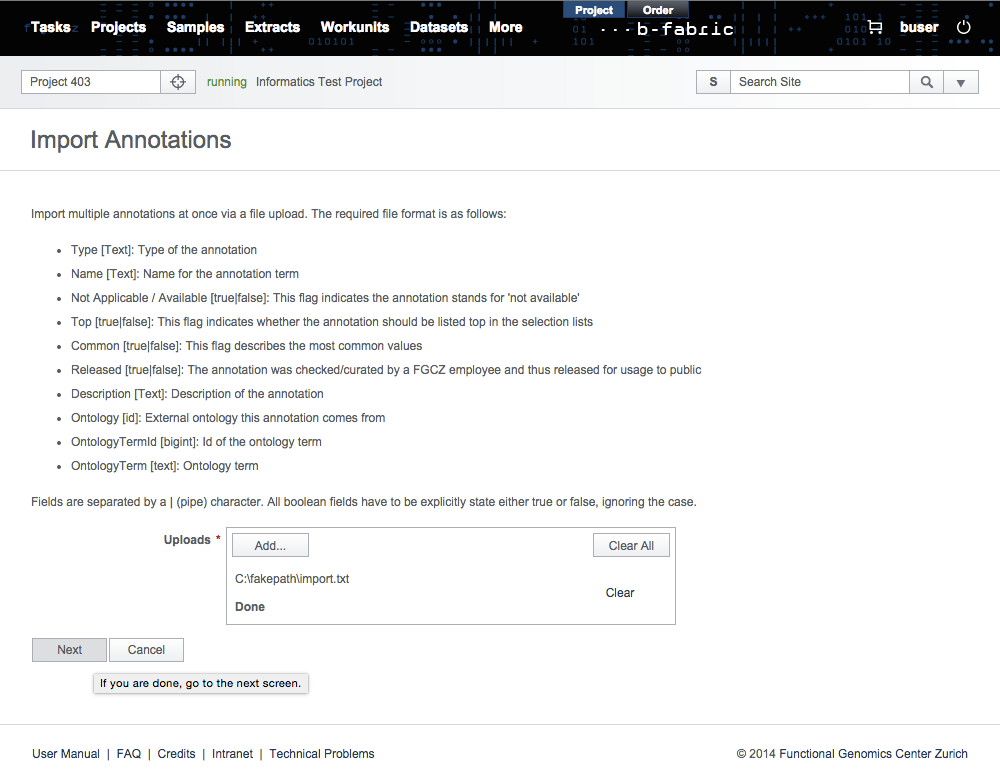
Import multiple annotations at once via a file upload. The required file format is as follows:- Type (Text): Type of the annotation
- Name (Text): Name for the annotation term
- Not Applicable / Available (Boolean): This flag indicates the annotation stands for 'not available'
- Top (Boolean): This flag indicates whether the annotation should be listed top in the selection lists
- Common (Boolean): This flag describes the most common values
- Released (Boolean): The annotation was checked/curated by an FGCZ employee and thus released for usage to public
- Description (Boolean): Description of the annotation
- Ontology (Id): External ontology this annotation comes from
- OntologyTermId (Bigint): Id of the ontology term
- OntologyTerm (Text): Ontology term
Fields are separated by a "|" (pipe) character. All boolean fields have to be explicitly state either true or false, ignoring the case.
Assume that the following file import.txt contains the annotations to be imported locally:
import.txt
Species | Toxoplasma Gondii | false | false | true | false | Tiger isolate || Strain | Tigerisolat | false| false | true |false | Oozysten aus Tiger isoliert, mit grosser Wahrscheinlichkeit Typ2 ||
Click on the Import Annotations via File link in the Admin section and select import.txt.
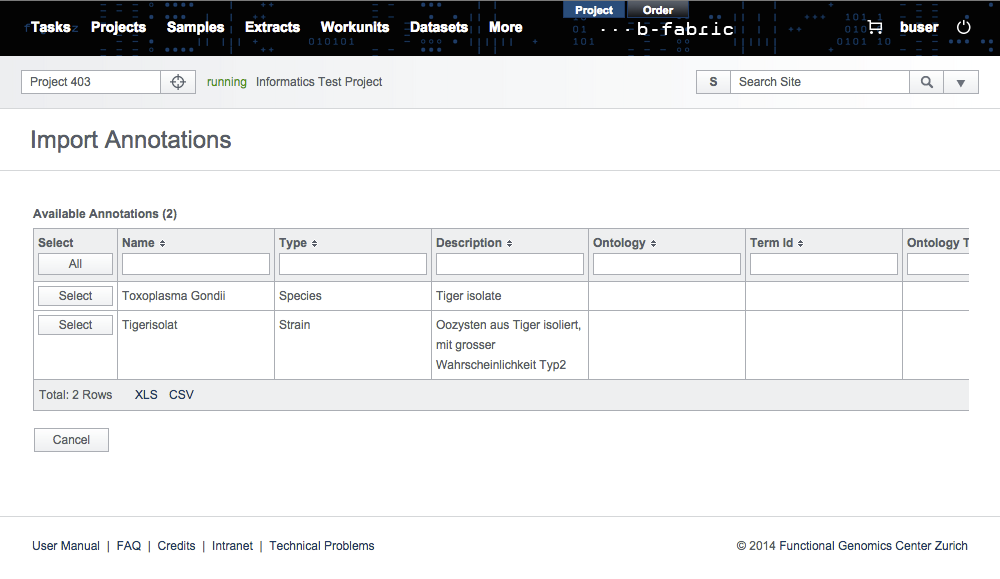
After clicking on the Next button, all the valid annotation in the import file will be loaded into the form on the screen. There you can still select all or specific annotations to import.
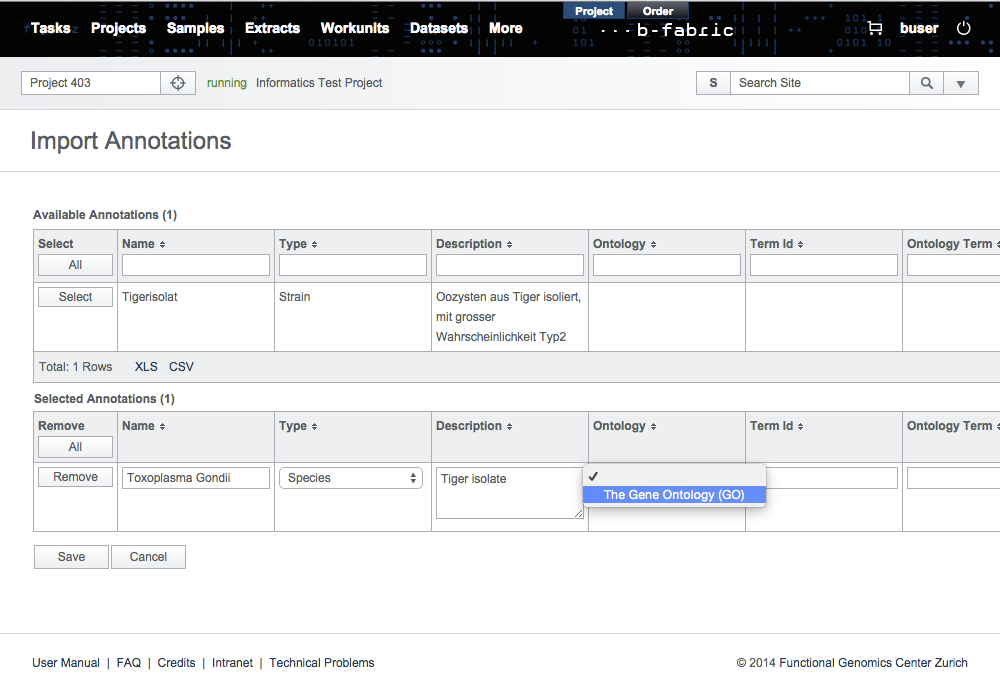
The next step is to set the types of imported annotations manually. This is a required step since annotation types differ from ontology to ontology. Then, click on Save to store the annotations in the B-Fabric vocabulary.
Import Annotations from External Ontologies
When importing annotations from external ontologies, there are two different cases. The first case is to directly import annotations from an external ontology. This operation is performed by a user with the annotationManager role and often used to add several annotations at once to the B-Fabric vocabulary. The second case occurs spontaneously. For instance, when a user wants to add an annotation that does not exist in B-Fabric’s vocabulary while creating a new sample.
Import Annotations Directly into B-Fabric vocabulary
Annotations from external ontologies can be imported directly by clicking on the Import Annotations via Search link in the Admin section.
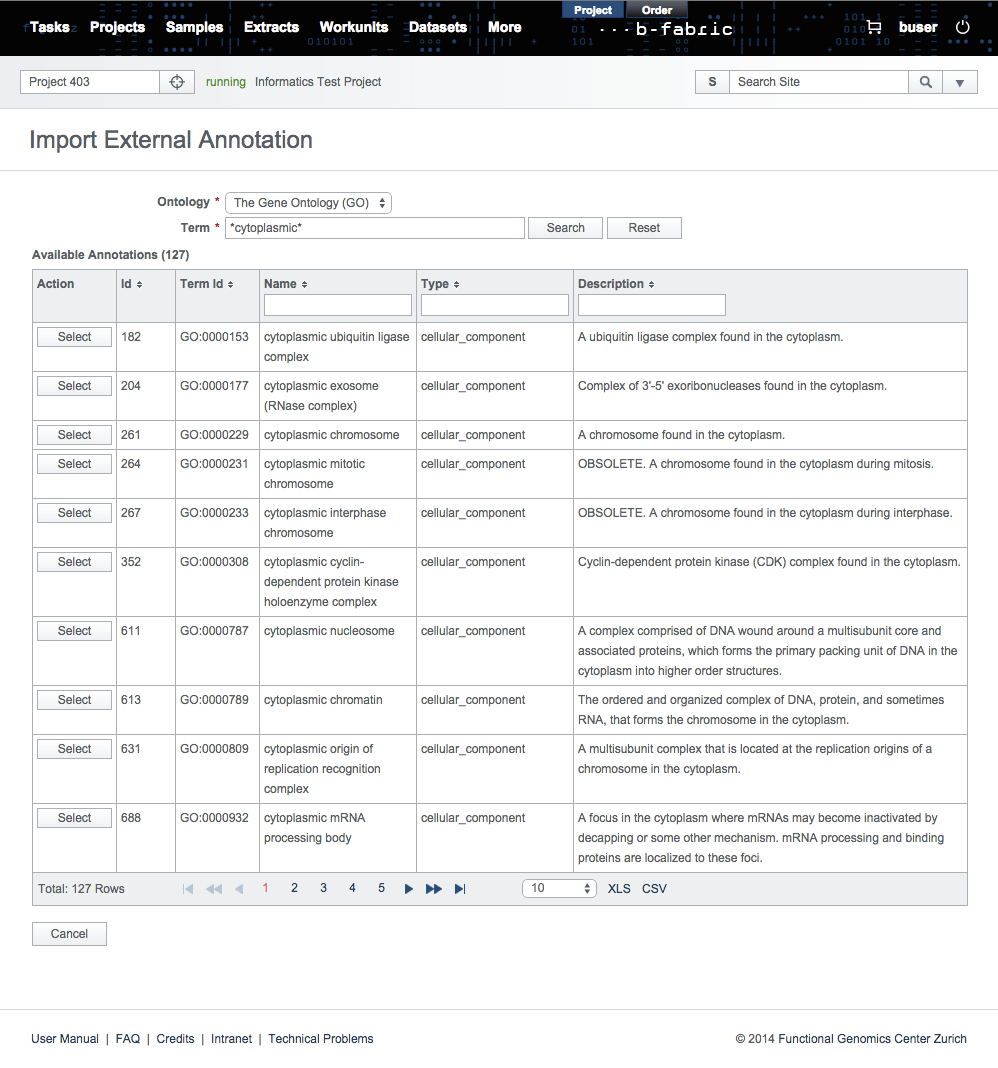
In the search form, select the ontology and enter your search term. Ontology, currently, is only bound to the Gene Ontology (GO). The search term can contain wildcard characters "%" and "_".
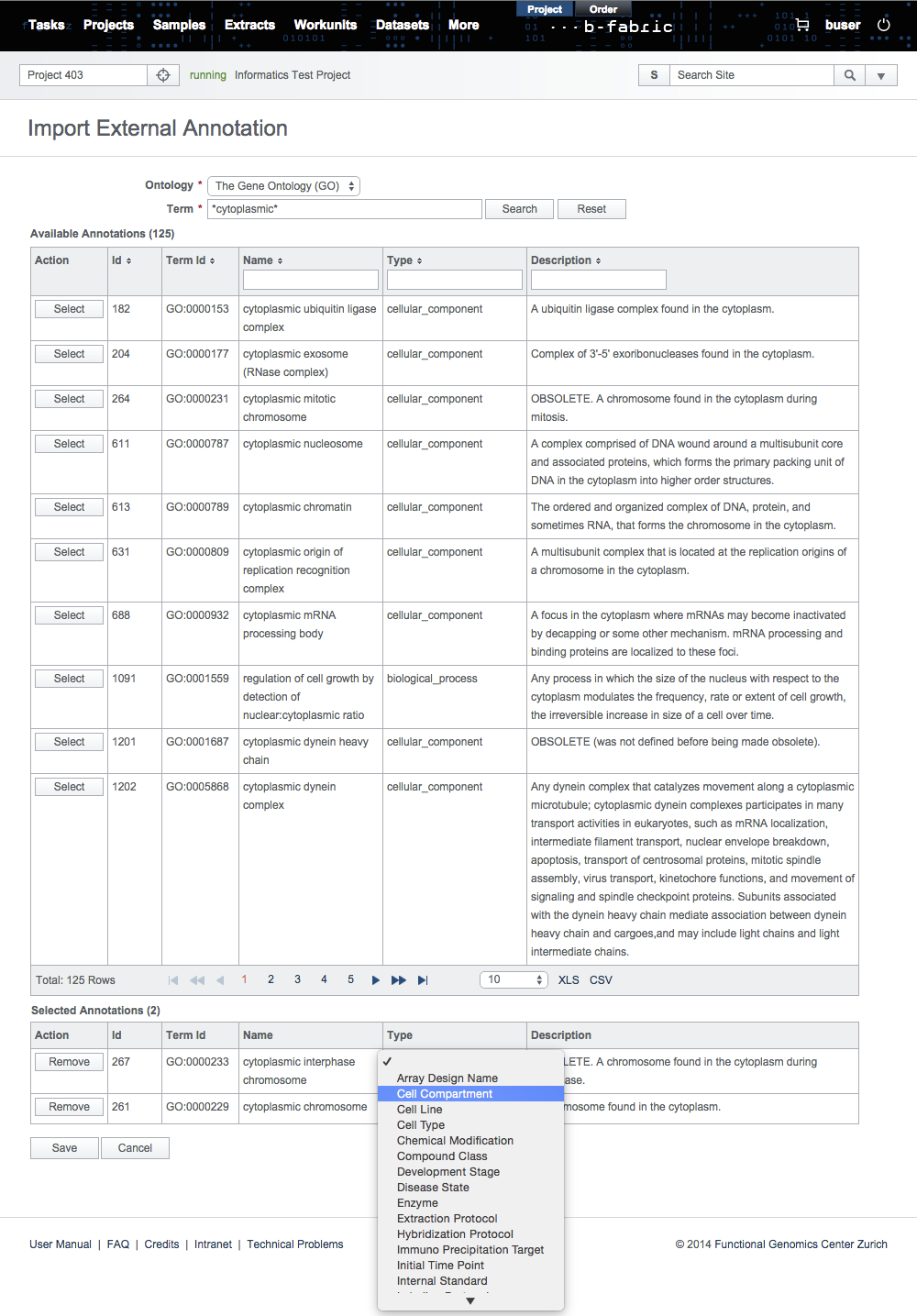
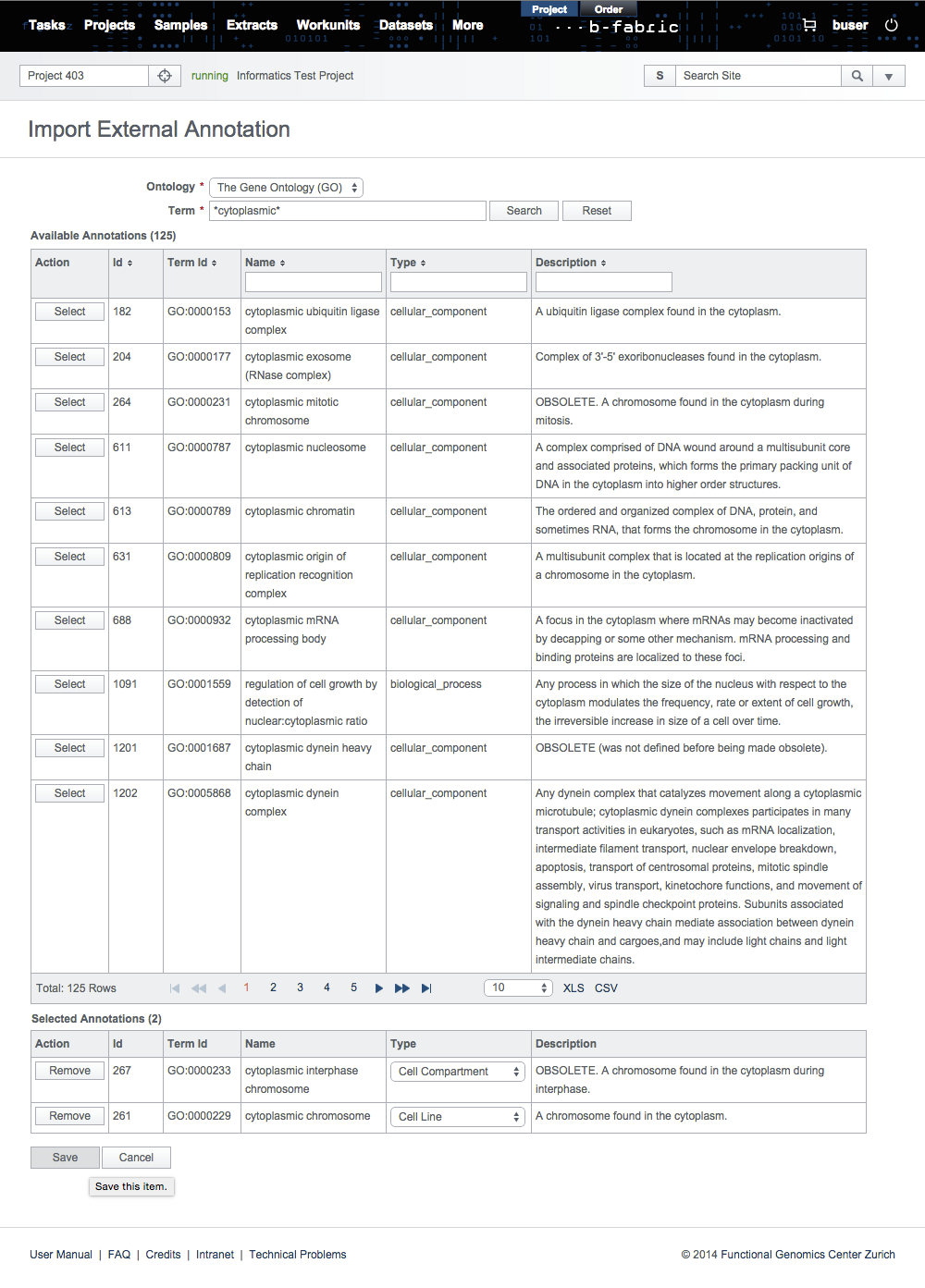
In the search results, select the annotations to be imported and set their annotation types manually.
Then, click on Save to store the annotations in the B-Fabric vocabulary.
Import Annotations Indirectly into B-Fabric vocabulary
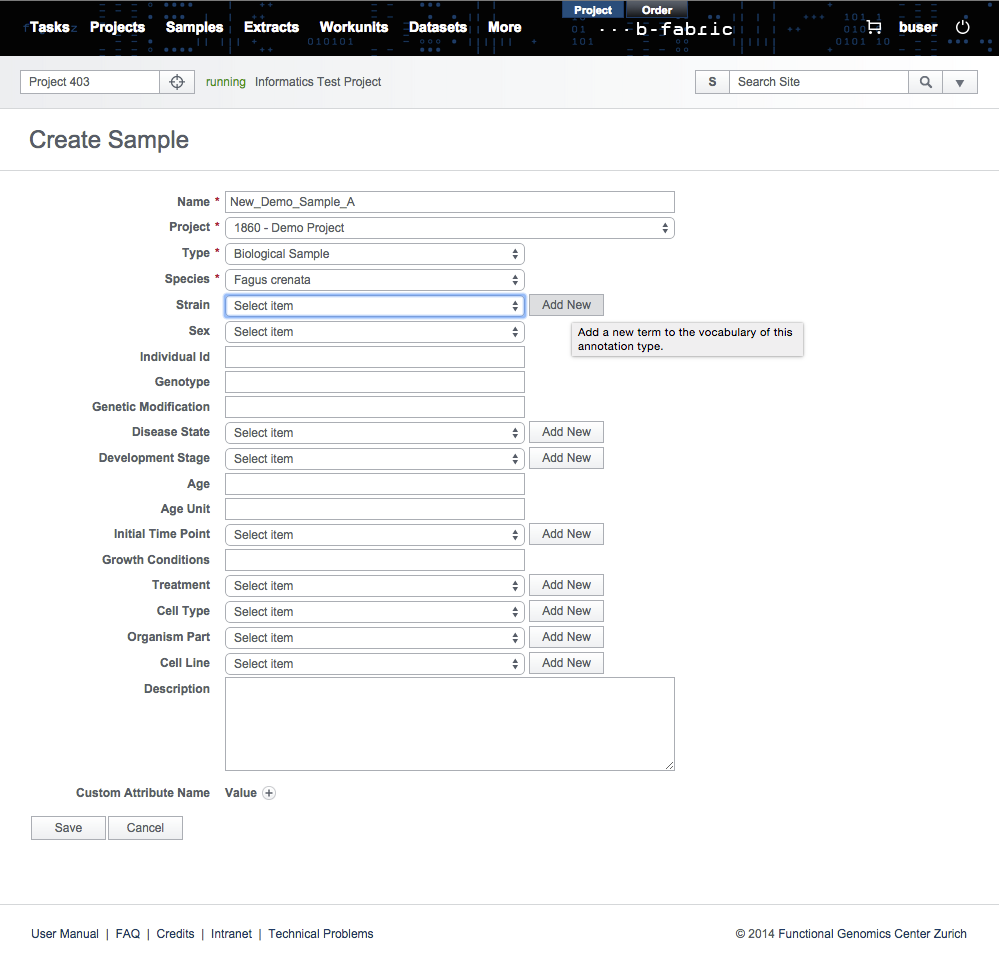
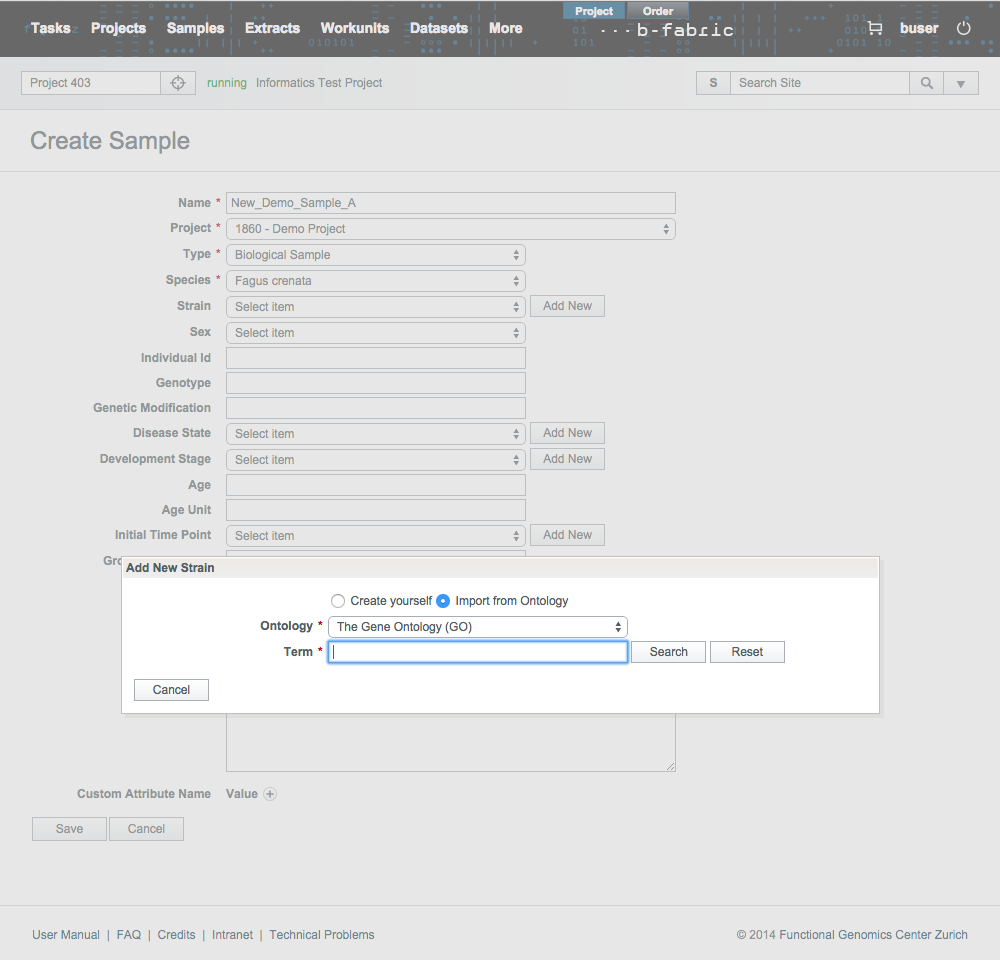
In several data entry forms provided by B-Fabric, it is possible to extend the B-Fabric vocabulary indirectly. For instance, while creating a new sample, if you can not find the strain you are looking for in the drop down list, you can add your new strain by simply clicking on the Add New button seen in the picture below. Pressing this button means that the user wants to select a single annotation from some ontology and to add it to the B-Fabric vocabulary.Clicking on Add New button causes a pop up form to appear. This form will allow the user to create an annotation manually or to search over some ontology for a specific term and, if found, to add it to the B-Fabric vocabulary. Click on ''Import from Ontology" radio button to select the new annotation from an existing ontology.
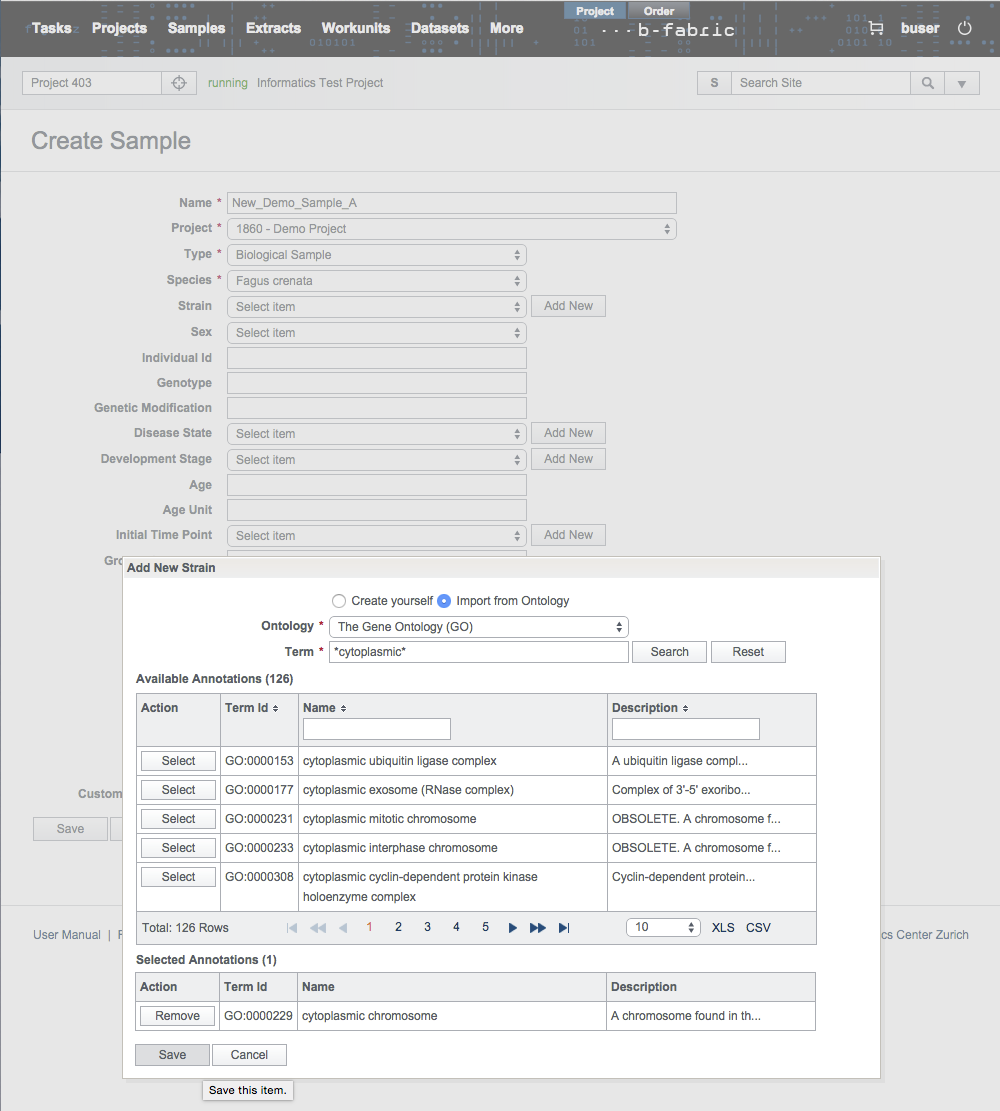
Fill in the search form accordingly.
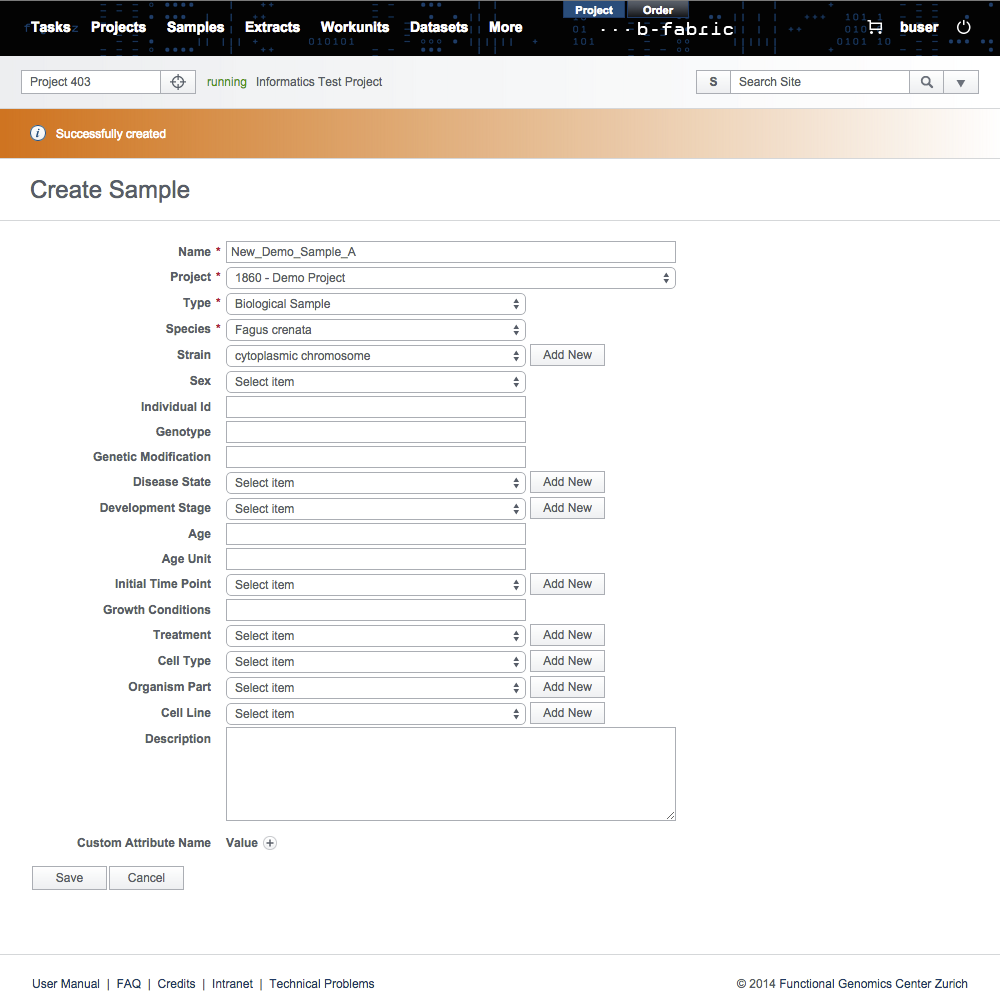
In the results, you can select only a single annotation and save it in the B-Fabric vocabulary.
Handling Complex Annotated Objects
Often, users want to reuse and adapt existing sample and extract information in other projects. In addition to copy and move single samples and extracts, users demand for copying and moving entire sample and extract hierarchies. To achieve these, B-Fabric supports the following copy and move functionalities:- Shallow Sample Copy: Copy only a single sample object without any extracts derived from this sample.
- Deep Sample Copy: Copy a single sample object together with the entire extract object hierarchy (transitively) derived from this sample object.
- Shallow Extract Copy: Copy an extract object together with all its ancestors, i.e., the parent samples and parent extracts.
- Deep Extract Copy: Copy an extract object together with its ancestors, i.e., its parent samples and parent extracts, and all its descendants, i.e., derived child extracts.
- Sample Move: Move a sample to a project together with its entire extract hierarchy. Copy all ancestors of the involved extracts in the sample's hierarchy to the new project.
- Extract Move: Move an extract together with its entire child extract hierarchy. Copy all ancestors of the involved extracts to the new project.
Note that extracts can be bound to resources. This is the case, after a workunit has been created and its resources have been assigned to extracts. In such cases, an extract move cannot be supported without requiring the movement of the underlying resources.